Molecular dynamic simulations reveal detailed spike-ACE2 interactions
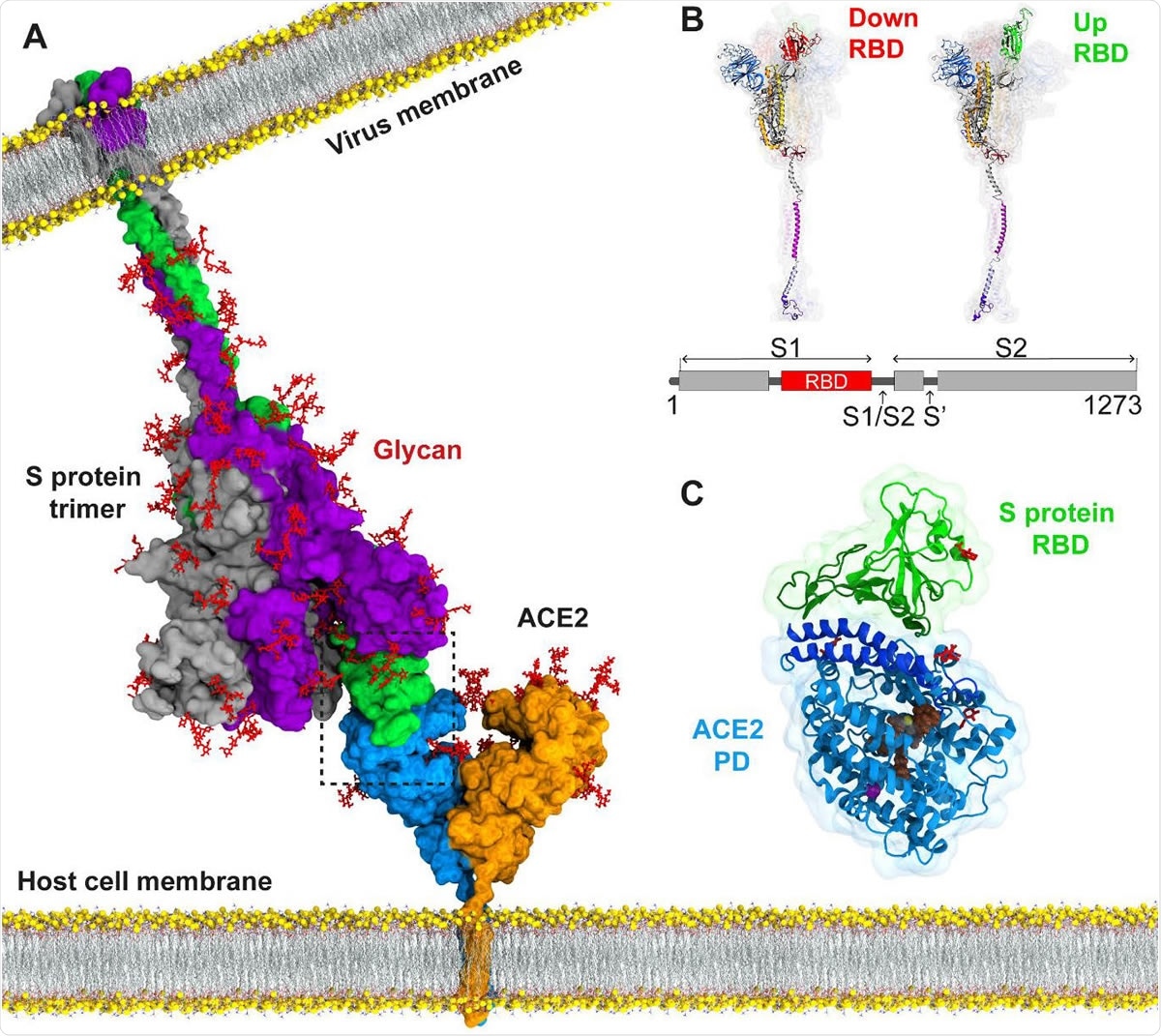
The current COVID-19 pandemic has spread throughout the world. Caused by a single-stranded RNA betacoronavirus, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which is closely related to but much more infectious than the earlier highly pathogenic betacoronaviruses SARS and MERS-CoV, has impacted social, economic, and physical health to an unimaginable extent.

Repurposing Therapeutics for COVID-19: Supercomputer-Based Docking to the SARS-CoV-2 Viral Spike Protein and Viral Spike Protein-Human ACE2 Interface - Abstract - Europe PMC
An experimental peptide could block Covid-19, MIT News

Biology, Free Full-Text
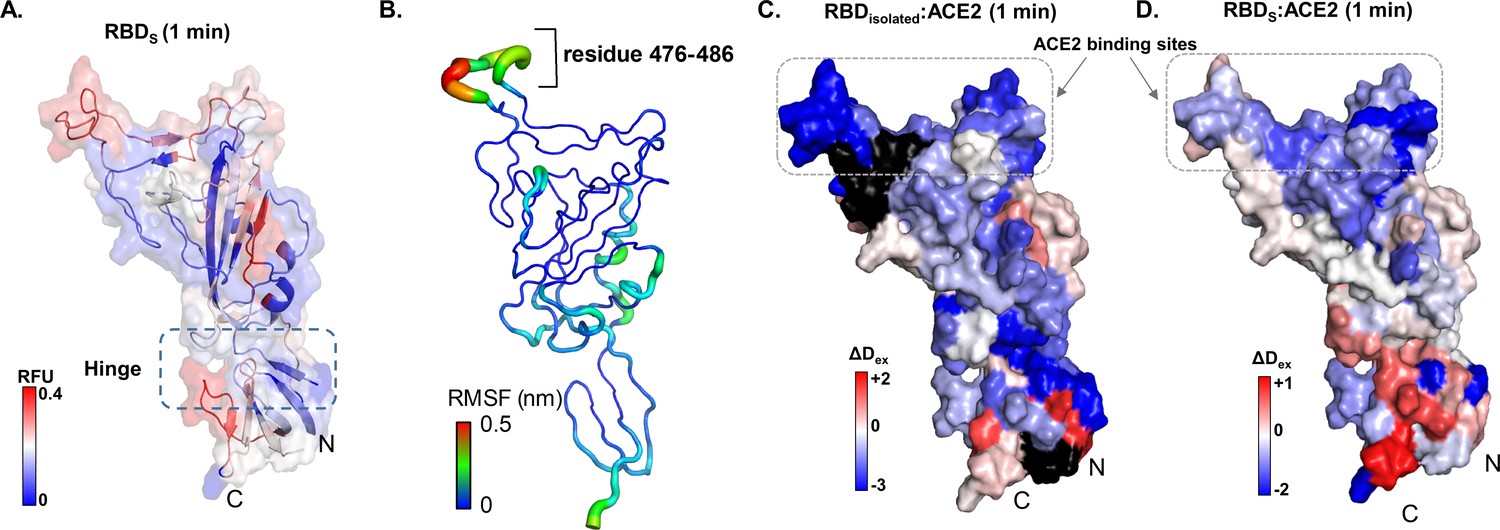
SARS-CoV-2 S protein:ACE2 interaction reveals novel allosteric targets

Binding affinity between coronavirus spike protein and human ACE2 receptor - Computational and Structural Biotechnology Journal
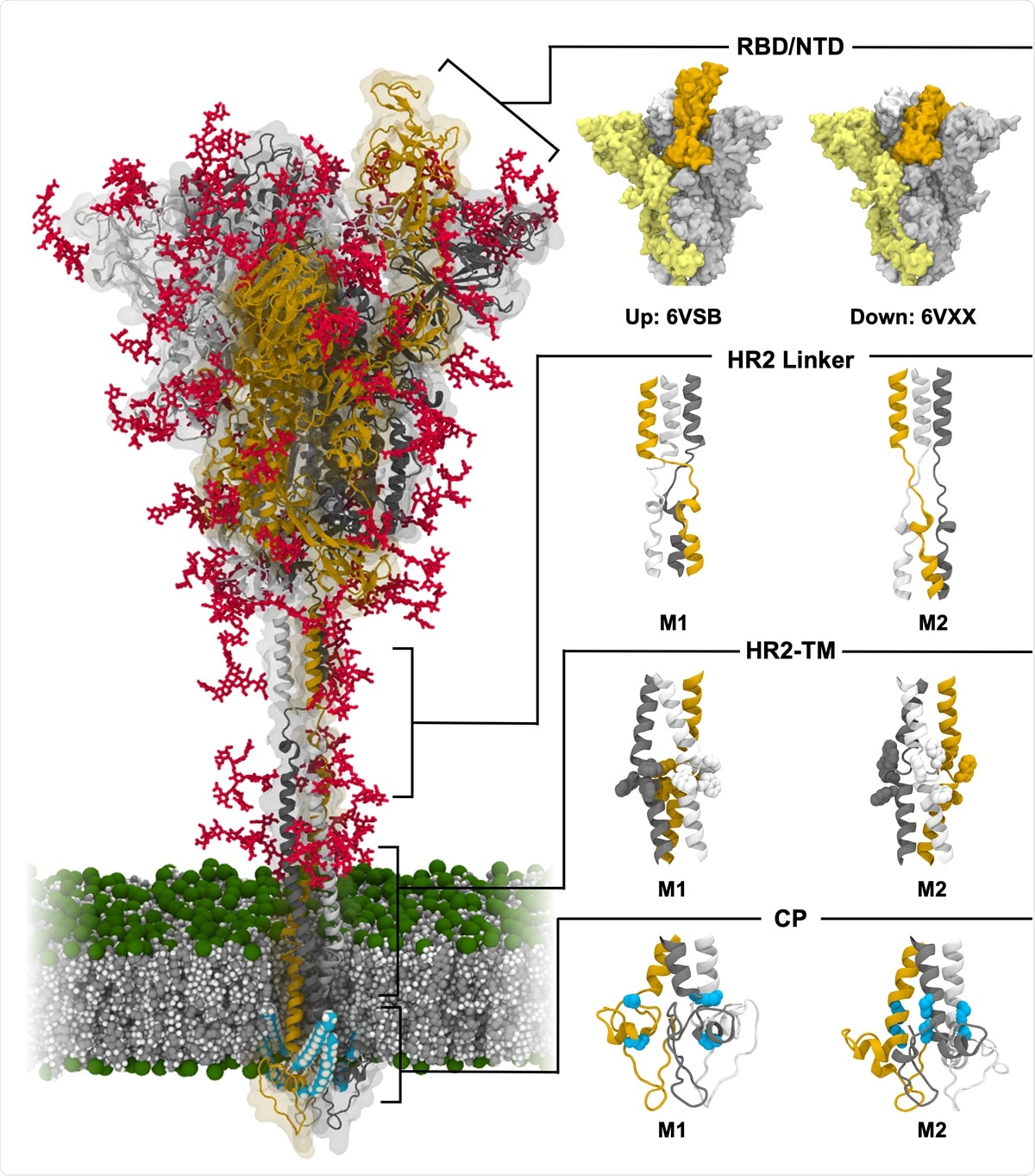
Molecular simulations reveal detailed structure and dynamics of SARS-CoV-2 spike protein

Full article: Virtual screening and molecular dynamics simulation study of abyssomicins as potential inhibitors of COVID‐19 virus main protease and spike protein

Molecular dynamic simulations reveal detailed spike-ACE2 interactions

Cells, Free Full-Text
Molecules, -Text
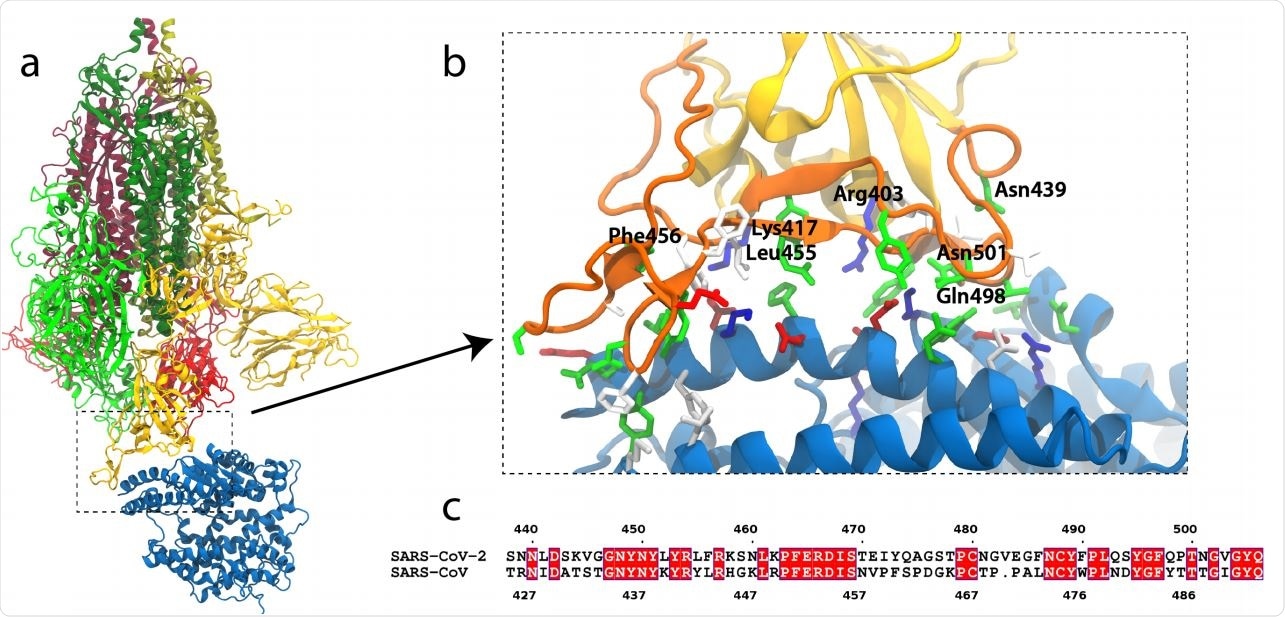
Ai helps identify critical interactions for SARS-CoV-2 spike protein binding to ACE2
Unraveling SARS-Cov-2 spike protein conformational dynamics under the influence of electric fields - NHR4CES

Mutational landscape and in silico structure models of SARS-CoV-2 spike receptor binding domain reveal key molecular determinants for virus-host interaction, BMC Molecular and Cell Biology